RNA Technologies aims to study RNA to fundamentally understand biology, and so to find druggable pockets in the molecular processes of cells. Bioinformatics and computational tools in general are already the hallmark of IIT’s research on RNA. Several research pipelines have been implemented and are used daily at IIT. RNA therapeutics will take advantage of the studies developed at CMP3VdA and the Joint Labs with several clinical partners as the Gaslini Children’s Hospital and the San Martino Hospital in Genoa.
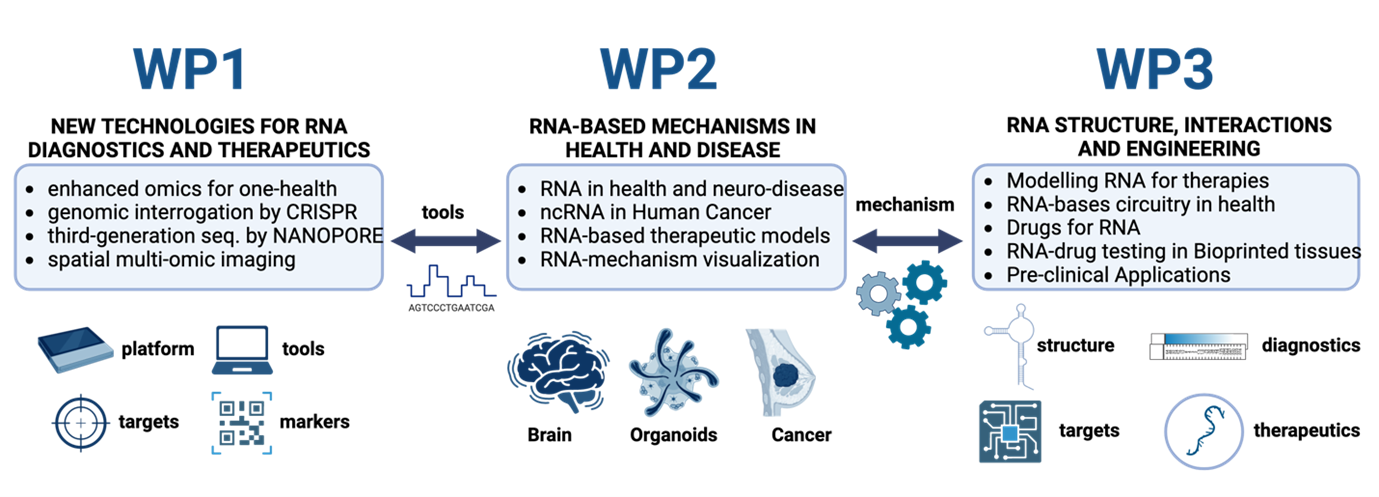
The RNA Technologies Flagship pursues interconnected objectives to transform RNA biology into actionable therapies and
diagnostics. It drives revolutionary characterization using advanced sequencing, CRISPR tools, and omics technologies for deep insights into RNA mechanisms, particularly in neurological diseases, cancer, and health contexts. Simultaneously, it advances therapeutic modeling through computational predictions, engineered RNA devices and drug discovery to unlock precise, safe interventions.
The ambition of the RNA Technologies Program is to drug the undruggable.