Technologies
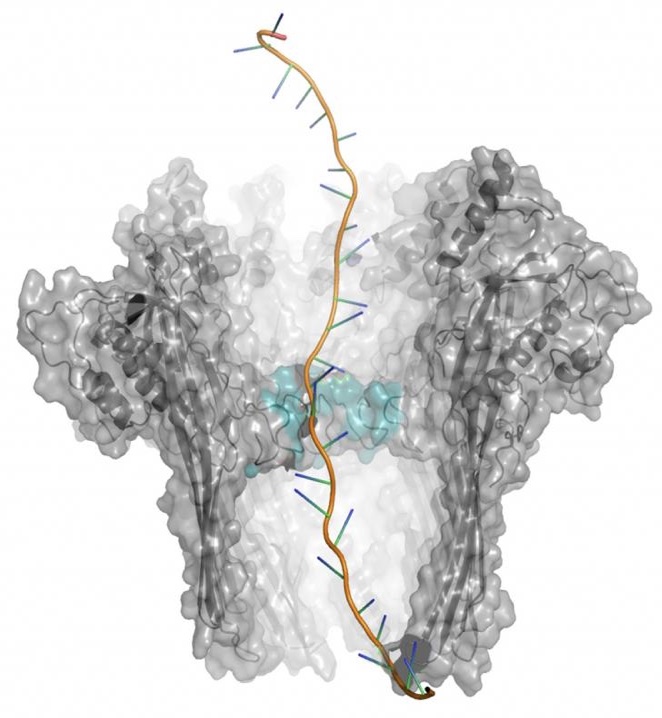 |
Single-molecule sequencing (Nanopore)Long-read and direct RNA sequencing are used to resolve transcript structure at full-length resolution, enabling the identification of isoforms, non-coding RNAs, and RNA modifications within the same experimental framework. These approaches support transcript-resolved analyses across cell lines, organoids, and clinical samples, forming the basis for integrated transcriptomic and epitranscriptomic profiling. |
|
|
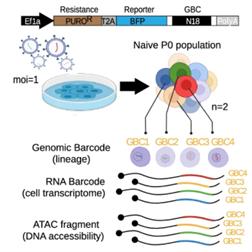
Clonal tracking and lineage tracingGenetic and transcript-based barcoding strategies are used to track cell populations over time, linking clonal dynamics with transcriptional states. These approaches enable the study of tumour evolution, treatment response, and the emergence of specific cellular programs in heterogeneous systems, including single-cell contexts. Within the GALILEO framework (Nadalin et al. 2024), clonal tracking is directly coupled with single-cell multi-omic profiling, enabling the simultaneous reconstruction of lineage, transcriptional states, and chromatin accessibility at single-cell resolution. |
Clonal tracking and lineage tracingGenetic and transcript-based barcoding strategies are used to track cell populations over time, linking clonal dynamics with transcriptional states. These approaches enable the study of tumour evolution, treatment response, and the emergence of specific cellular programs in heterogeneous systems, including single-cell contexts. Within the GALILEO framework (Nadalin et al. 2024), clonal tracking is directly coupled with single-cell multi-omic profiling, enabling the simultaneous reconstruction of lineage, transcriptional states, and chromatin accessibility at single-cell resolution. |
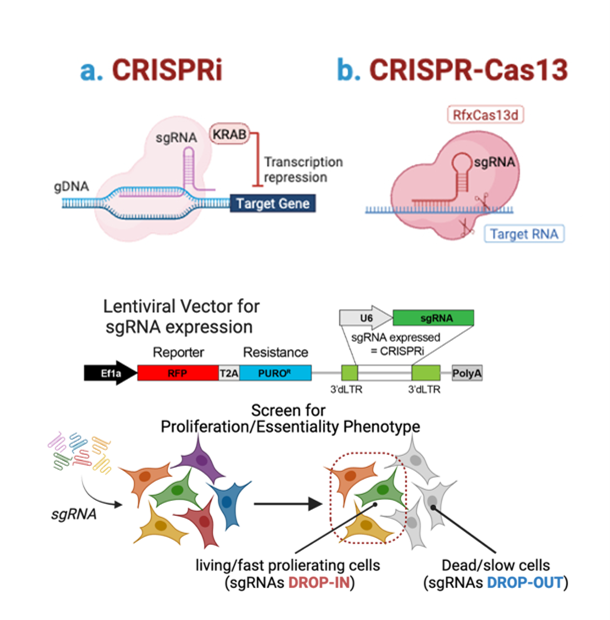 |
CRISPR functional genomicsCRISPR interference (CRISPRi) platforms are used for systematic perturbation of coding genes, non-coding RNAs, and regulatory elements. Pooled screening approaches enable the identification of functional dependencies and regulatory vulnerabilities, while integration with transcriptomic readouts allows the dissection of underlying gene regulatory networks. |
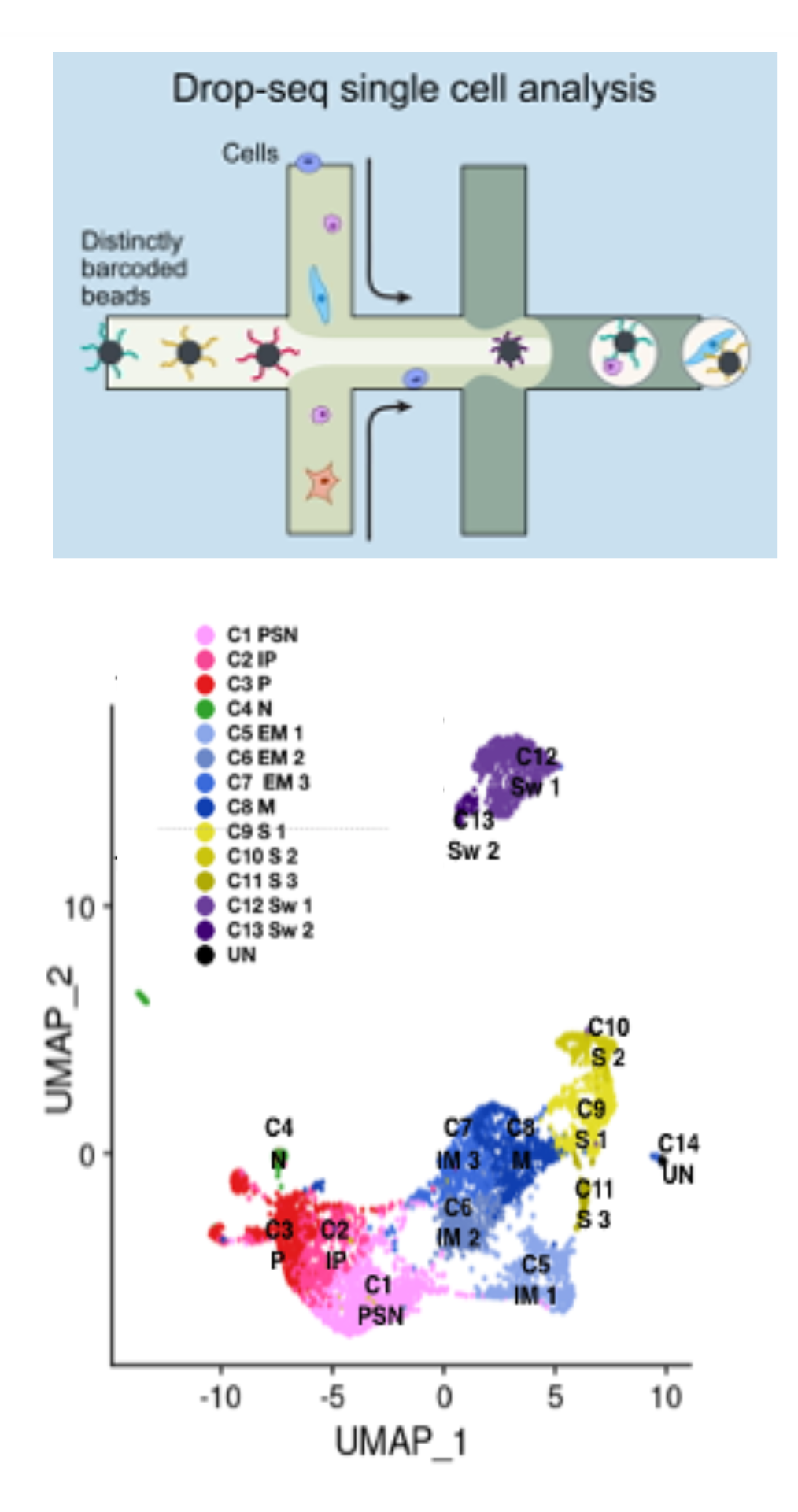 |
Single-cell and multi-omics genomicsSingle-cell RNA-seq and multi-omic approaches are used to profile gene expression and chromatin accessibility at high resolution. These technologies enable the identification of cellular subpopulations, regulatory programs, and dynamic states, and can be integrated with perturbation and lineage tracing strategies to reconstruct functional cellular trajectories. |